The Human Genome is a Mess
This week's Nature has a News Feature entitled "Human genome: patchwork people" that focuses on the amount of structural polymorphisms (rearrangements, segmental duplications, large deletions) polymorphic in human populations:
"But even as the ink was drying on the complete [human genome] sequence, some researchers were questioning whether there was really such a thing as the definitive edition of the Book of Life. By skim-reading individual genomes, these scientists were finding bizarre and unexpected irregularities. In some people, whole paragraphs of the text were duplicated, whereas in others, large passages were missing, or even printed backwards."Gee wiz, who would have thought humans would have structural polymorphisms segregating in natural populations!? I mean, it's not like Drosophila geneticists haven't known about these things for seventy years. But golly, we thought these things caused diseases in humans because we're so gosh darn (ok, I've gotta stop using the stupid 1950s exclamations) special:
"These major revisions turned up in all kinds of people, including many who seemed healthy and normal."Um, that's because of ascertainment bias. You see, when you only screen for big fuck ups in the genome of sick people, you're only gonna find screwed up genomes in, well, sick people. Guess what, we've know for a while that "healthy" folks have "abnormal" genomes.
In case the jargon was too much, about half of the "anomalies" caused "clinically significant" problems -- that means half of them did not. Many of the abnormalities described resulted in aneuploid individuals (those with an extra copy or missing a single copy of a single chromosome, ie, monosomic or trisomic), and these are probably not heritable. Many of them, however, had inversions or translocations, which are heritable and can increase in frequency in a population due to drift or selection."The results of six major surveys of consecutive newborns show that 6 out of 1000 (0.6%) carry one or another kind of chromosome anomaly. The breakdown is as follows: 0.22%, sex-chromosomal abnormalities; 0.14%, autosomal trisomies (+D, +E and +G); 0.19%, balanced structural rearrangements and 0.06%, unbalanced structural rearrangements and others. The frequency of "clinically significant" anomalies has been estimated to be about one-half of the total of all chromosomal abnormalities detected in newborns" (Sankaranarayanan 1979).
We've known about the Drosophila inversion polymorphisms for so long because of polytene chromosomes. We can view the chromosomes by extracting the salivary glands out of larval flies where a single cell has multiple copies of each chromosome stacked on top of each other. In individuals heterozygous for large deletions or inversions, these appear as loops in the polytene chromosomes.

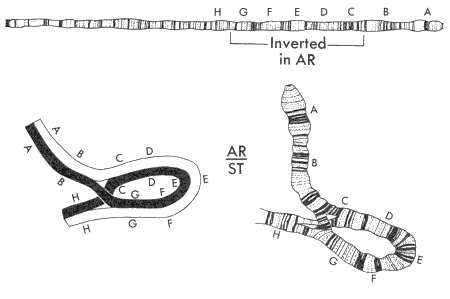
We cannot do this in humans, but now that we have whole genome sequences, there are neat bioinformatic ways to detect large scale chromosomal rearrangements. For instance, some parts of the genome seem to get sequenced twice as often in whole genome sequencing projects -- this is indicative of recent large scale duplications. We can also use single nucleotide polymorphism (SNP) data to detect regions of the genome with abnormal levels of polymorphisms, suggesting some sort of large scale rearrangement.
It's good to see that mammalogists are beginning to examine the importance of these rearrangements outside of their detrimental qualities."Genome researchers now have a catch-all phrase for the vast array of rearrangements -- including copy-number polymorphisms, inversions, deletions and duplications -- that occur normally in the human genome. They call it structural variation, and have described at least 800 individual variants that, in total, account for about 3.5% of the human genome
. . .
"What's more, statistical analyses show that regions of structural variation contain genes that are still evolving in humans. If these genes are important enough for evolution to be changing them, they must affect us in some way, for better or worse."
Check, E. 2005. Human genome: patchwork people. Nature. 437: 1084-1086.
Sankaranarayanan, K. 1979. The role of non-disjunction in aneuploidy in man an overview. Mut. Res. 61: 1-28.









0 Comments:
Post a Comment
<< Home